Finding the Best DNA Health Test
Our experts explain everything you need to know about DNA health testing.
Even though direct-to-consumer genetic testing has existed for about a decade, beginning with the National Geographic Society and IBM’s launch of the Genographic Project in 2005,1 scientists only finished the first complete sequence of a human genome in 2022.2 With such huge advances in our understanding of genetics and what they could mean for our health, it’s no wonder that there are so many DNA health tests on the market today.
But with so many options, it can feel overwhelming trying to figure out the right test for you. Maybe you want to learn about all the nitty-gritty details that make up your genome, or you could be interested in uncovering your carrier status, or perhaps you’re more interested in finding out ways to improve your health based on your results — no matter your reasons for wanting a DNA health test, there’s at least one out there to suit your needs.
In this guide, we’ll cover all of the pros, cons, and details of our top three DNA health tests (along with a couple of honorable mentions) to help you find the ideal testing service and platform.
If you’re in a hurry, here’s a quick summary of our findings.
Summary of recommendations
- Best overall: Nebula Genomics
- Best budget pick: MyToolbox Genomics
- Best for nutrition and fitness: InsideTracker
- Best for carrier screening: Nebula Genomics
- Best epigenetic test: MyToolbox Genomics
- Best full genome test: Nebula Genomics
- Fastest turnaround time: InsideTracker
Nebula Genomics’ whole-genome DNA testing helps you learn a great deal about your genetic mutations, carrier status, traits, and more.
With interactive tools and weekly updates to your results, whole-genome sequencing from Nebula Genomics offers a wealth of up-to-date information and valuable insights into how your DNA may be impacting your health. On top of that, you’ll also receive ancestry and oral microbiome data. And compared to competitors, these tests are relatively affordable (especially during one of Nebula’s frequent sales).
With MyToolbox Genomics, combining DNA and epigenetics results can help you learn about how your lifestyle and environment may be impacting your genes — and your health.
MyToolbox Genomics offers DNA and epigenetics test kits to help you learn about what’s set in stone with your genes and what you can potentially change for the better. Epigenetics test results come with personalized guidance (such as diet and exercise recommendations) to help you along the way. A subscription plan for epigenetics testing lets you save money while retesting every three, six, or 12 months to see how your lifestyle changes are working.
In this Review
Why you should trust us
Over the past two decades, Innerbody Research has helped tens of millions of readers make more informed decisions to live healthier lifestyles.
We extensively research and test each service and product we review, including each of these DNA tests. All told, our team has spent 300+ hours testing and researching DNA testing services to provide you with details and information that can only be found through hands-on experience with these products. We read dozens of scientific studies on DNA testing, and our group of testers purchased and completed the DNA health tests, then documented their personal thoughts and experiences with things like wait times, customer service, reading results, and more.
Additionally, like all health-related content on this website, this guide was thoroughly vetted by one or more members of our Medical Review Board for accuracy.
Top considerations and evaluation criteria
A lot of thought goes into figuring out which DNA test is best for learning about your desired health information. The process ultimately involves a fair bit of investment and trust — you’re investing time and money into purchasing a test from a company to which you’re entrusting personal information in hopes that you’ll receive accurate and helpful results.
With this in mind, we narrowed down our evaluation criteria to include five main points:
- Accuracy
- Cost
- Privacy
- Actionability
- Speed
Accuracy
Winner: Nebula Genomics
Unless you conduct a bit of research beforehand or have some prior education on the topic, It can be hard to tell whether or not a DNA test is accurate. On top of that, what your DNA says isn’t a perfect parallel to your lived experience — just because you’re genetically predisposed to something doesn’t mean that you’ll experience it or develop in that way.3 For instance, someone might have genes that suggest a short stature, but they’re actually average height or even tall. But how can you tell the difference between this type of circumstance and a test that’s just plain incorrect?
Every analyzed sequence provides an opportunity for mistakes — even current next-generation sequencing (NGS) technologies, while highly accurate in their own way,4 also have an estimated error rate “between 1 per 100 and 1 per 1,000 base pairs sequenced.”5
Our choice for the best overall DNA health test, Nebula Genomics, is a full genome test (meaning it analyzes each and every one of your genes). The Deep and Ultra Deep tests, in particular, read through and check your entire genome at least 30 and 100 times, respectively. Competitor Sequencing.com is another full genome test that checks samples at least 30 times, but that’s the only depth offered in its tests. 30× sequencing coverage is recommended by researchers for human genomes, and it’s typically all that’s necessary to detect most common and rare variants.6 However, under specific circumstances — like if you’re looking for an exceptionally rare mutation — then opting for Nebula’s Ultra Deep test with 100× coverage may be worthwhile.
Nebula Genomics also offers a vast collection of materials to help you learn about your genome, including guides, studies, genome exploration tools, and more. The company is highly transparent in what research is used to come to each result or conclusion.
Our other recommendations also boast good scientific accuracy. MyToolbox Genomics, for instance, isn’t a whole genome test (it only checks certain genes), but it does back up your results with a host of studies. And InsideTracker boasts a peer-reviewed scientific study that suggests following its Action Plans may lead to positive outcomes.7
Cost
Winner: MyToolbox Genomics
While the cost of a DNA test has shrunk dramatically over the last few decades — going from hundreds of thousands (or millions for a full genome sequence) to hundreds of dollars — they still aren’t inexpensive.8
With that in mind, we compared the flat costs of the tests themselves (some companies offer multiple) and membership options where applicable. Some companies, like Nebula Genomics and 23andMe+, require a monthly or annual membership fee to maintain access to your results and information. We also considered exactly what you’re getting for the price — whether it’s a genetic trait test, full genome sequencing, or epigenetics test.
MyToolbox Genomics has no subscription fee and offers some of the least expensive costs overall. The company’s most expensive package, which contains both a genetic and epigenetic test, costs $299 and doesn’t require a membership or subscription. A close runner-up for this price is Nebula Genomics’ Standard kit with a lifetime membership; this option will also run you $299, but it doesn’t check for genetic mutations like the Deep and Ultra Deep options. Because of this, MyToolbox Genomics wins out overall for cost.
However, no company is perfect, and MyToolbox Genomics is no exception. While you won’t be charged membership fees, the company does suggest you retest your epigenetics every few months. If you’re interested in seeing how MyToolbox’s personalized suggestions (for things like vitamins, nutrition, and exercise) are working for you, then utilizing the company’s subscription service could save you money over time. Even testing once per year on the subscription plan takes the price of an epigenetics test from $273 down to $216. Keep in mind that retesting isn’t required, nor will you lose any benefits or bonuses by not participating — it’s simply a way to check up on how your lifestyle changes are working.
Privacy
Winner: Nebula Genomics
When it comes to your genes, privacy is key. Your exact DNA sequence can only be found in you, but traces can be found in your family by blood. It can be easy for hackers to learn sensitive information about you and your family if there are any data breaches, even if your family hasn’t taken part in a DNA test. As a recent example, 23andMe suffered from a pretty massive data breach — almost seven million users’ ancestry data (and some health information) was compromised by an anonymous hacker.9
No company has the perfect privacy policy and security measures. However, Nebula Genomics stands out as the best choice for strong privacy.
Besides limiting access to third parties and stripping your data of identifying information, Nebula has also partnered with Oasis Labs to integrate a security product called Parcel into the company’s platform. This tool can help secure client data and create tamper-proof logs of data access history. In addition to Parcel, Nebula also lists three different cybersecurity companies employed to protect various parts of the platform — from checking for vulnerabilities and threats to improving cloud security and implementing quick fixes in the event something goes wrong.
MyToolbox Genomics has a slightly more middle-of-the-road approach to privacy. While it doesn’t boast the high-end security technology of Nebula Genomics, your information is kept in-house; your sample is processed by a single partnered laboratory. The company’s privacy policy notes that your information is de-identified and anonymized. And, just like Nebula and InsideTracker, MyToolbox won’t sell your information to third parties, including researchers, without your consent.
One of our honorable mentions, Sequencing.com, also appears to care a lot about data protection — the company’s privacy policy is even dubbed “Privacy First.” This policy explicitly states that Sequencing.com will never sell or monetize your data, and you have the option to delete your account and information whenever you want. However, there is a pretty big drawback to this service. Since Sequencing.com is an app-based marketplace that uses many third-party applications to generate various results reports, you can’t know exactly what those third parties do with your genetic information. This is one of the biggest reasons why we prefer it when DNA testing companies keep all of your data in-house or with partnered labs.
Actionability
Winner: InsideTracker
Though it can be beneficial to learn about every part of your genetic makeup through whole genome sequencing, not every piece of information that you learn will be valuable. This is especially true if you don’t know what to do with your newfound knowledge. Being aware of your genetic predisposition for a health condition is one thing, and knowing what to do to stay healthy is another.
Out of our top picks, InsideTracker offers the most thorough guide toward optimal health, known as an Action Plan. InsideTracker’s DNA testing plays an interesting role — it almost acts as a supplemental test for the company’s other health tests, which require blood draws every three months for biomarker analysis. These blood tests measure and track things like inflammation, blood count, and vitamin levels. Along with your genetic information, InsideTracker uses these biomarkers to create a personalized Action Plan — a guide with up to five small goals — for your health.
These plans use a straightforward step-by-step process to help guide you in making the changes necessary for your health goals. Each plan lasts 12 weeks (three months), and you can add 3-10 weekly actions. Some of these actions are meant to be completed daily, while others may only need to be done a few times per week.
It’s worth mentioning that InsideTracker’s DNA results are really only helpful if you tack on one of the blood tests. Receiving results that only include a handful of genetic markers with no other information considerably reduces InsideTracker's actionability.
MyToolbox Genomics is a great second option if you aren’t ready to have 4-6 vials of blood drawn at your local laboratory (potentially every three months). The action plans from these tests aren’t as robust, but they cast a much wider net to give you more genetic information on traits and predispositions that can affect your health and minor substitutions and suggestions to go along with each. You can’t customize your goals like with InsideTracker, but MyToolbox offers custom meal plans, a personalized training regimen, vitamin recommendations, and other genetic health insights without the need for bloodwork.
Speed
Winner: InsideTracker
When you’re waiting to hear back about the results of a test involving your health, some anxiety is to be expected.10 There are many factors that can impact the turnaround time for a DNA health test — shipping speeds, sequencing time, mail delays, test type, and more.
Based on our testers’ experiences and what the companies generally state, you can expect a genetic test (testing only specific genes; not the whole genome) to take about 3-5 weeks to process, while a full-genome test will take between 10-14 weeks.
When we compared each company’s estimated results time with what our testers actually experienced, InsideTracker was the fastest test by a long shot. Our testers, expecting their results to come back in 4-6 weeks, were surprised to get them in only ten days. No other DNA health testing company we tried came close.
The chart below compares each company’s estimated processing times with our testers’ results. Only one company, Sequencing.com, took longer than expected. But, even then, it only exceeded the time frame by one week.
| Estimated time | Actual time | |
|---|---|---|
| Nebula Genomics | 12-14 weeks | 14 weeks |
| MyToolbox Genomics | 3-4 weeks genetic; 8-10 weeks epigenetic (total) | 3 weeks genetic; 7 weeks epigenetic (total) |
| InsideTracker | 4-6 weeks | 10 days |
| Sequencing.com | 10-12 weeks | 13 weeks |
| 23andMe | 3-5 weeks | 5 weeks |
How our top recommendations compare
Choosing a DNA test can be complicated — the information you’re seeking from a test may be vastly different from what someone else is. Each testing company has its pros and cons, and several of them offer unique features or results.
We’ve put together a comparative chart so you can get a sense of what each testing company offers.
And below you’ll find another chart that breaks down how our picks compare in terms of cost. We delve further into pricing details, including any applicable membership costs, in each company’s dedicated section.
| Least expensive test | Most expensive test | Membership required? | |
|---|---|---|---|
| Nebula Genomics | $99 (Standard kit) | $899 (Ultra Deep kit) | Yes (for all kits) |
| MyToolbox Genomics | $199 (DNA test) | $299 (DNA + Epigenetics Test) | |
| InsideTracker | $249 (DNA kit) | $2,682 (bundle of four Ultimate blood tests) | |
| Sequencing.com | $399 (Ehlers-Danlos screening test) | $1,999 (Whole genome sequencing expedited) | Yes (a free membership option is available, but you won’t be able to access your whole genome analysis) |
| 23andMe | $109 (Ancestry) | $1,188 (23andMe+ Total Health) | Yes (for 23andMe+ options and to receive updates on lower tiers) |
Who is at-home DNA health testing for?
Direct-to-consumer DNA health tests are best suited for people who are curious about how their genetic makeup is impacting (or may eventually impact) their health. Some of these tests (such as those from InsideTracker) can also be helpful for athletes and those interested in using their results in ways that can benefit their fitness, nutrition, and overall health.
Comprehensive whole-genome testing, from companies like Nebula Genomics, can also provide some valuable insight into whether or not you’re a carrier — this is someone who “carries” a variant of a gene that’s been associated with a disease (or a trait). Carriers don’t show symptoms of the condition they carry the variant for, but there is the risk that it can be passed down to offspring.11
Who should look elsewhere?
While at-home DNA health testing can technically be for anyone who wants to try it, you may have some unique needs or goals that may be better served in other ways.
- Medical advice: The results from nearly all at-home DNA health tests can’t legally be considered medical advice. The only test with results that can be accepted as such is 23andMe, which received FDA approval in 2017.12 However, we still recommend that you take any worrisome results to a doctor for further evaluation and confirmatory testing.
- Specialty tests: If you’re seeking genetic testing for certain conditions, like cancer or congenital disabilities, seeing an in-person genetic counselor who specializes in the condition may be the best option. They can provide medical advice and help you to better understand any worrying results.
- Ancestry information: Even though some DNA health tests offer ancestry services, your results won’t be nearly as robust as those from a company fully dedicated to ancestry testing, such as AncestryDNA.
What types of DNA tests are there?
For the most part, DNA tests vary based on the scope (how much of your total genome it looks at) and what exactly is being analyzed. A majority of direct-to-consumer, or at-home, DNA tests that you can order without a doctor’s guidance or prescription fall into three categories.
Genetic testing
Genetic testing focuses on analyzing or investigating a selection of specific genes. The selection can vary widely from one company to the next, and you’ll only receive results about the genes the test checks for. MyToolbox Genomics, 23andMe, and InsideTracker utilize genetic testing.
Epigenetic testing
Epigenetic testing monitors chemical changes that essentially turn sections of your genetic code on and off. These changes influence how DNA gets read and coded and, in turn, change how your genes manifest.13 Some epigenetic changes can be reversible: for example, after you stop smoking, epigenetic changes caused by carcinogens in tobacco might reverse and turn off.14 Since epigenetics change over the course of your lifespan, they are often good indicators of aging.15 MyToolbox Genomics offers epigenetic testing on its own or paired with its DNA test for further insight into your genes.
Whole genome testing
A whole genome test, like those from Nebula Genomics, maps out every base pair on all 23 sets of chromosomes. No stone is left unturned — even parts of your DNA that don’t map to anything scientists know of yet. Whole genome tests often take a lot longer to process and require a more hands-on approach to understand since there’s a lot more information in your results. But having this wealth of information also means that you’re primed to learn more about yourself as additional research is conducted over time.
However, whole genome testing doesn’t include your epigenetics. If you’re looking for information on those chemical changes, you’ll need to take a specific test for them, such as the one offered by MyToolbox Genomics.
Other types of genetic tests are less common or only orderable by a genetic counselor or medical professional, such as:
- RNA testing
- Mitochondrial DNA testing (for maternal ancestry)
- Exome testing
- Chromosomal tests
- Karyotypes (which count and measure all of your chromosomes)
- Gene expression/mRNA tests
- Y-chromosome ancestry tests (for paternal ancestry)
For all of these tests, you’ll need to collect cells that contain complete sets of your DNA. Typically, DNA tests you take at home require samples of cells from your saliva (spitting into a tube) or from light brushing on the inside of your cheek with a swab.
What information can a DNA test tell me?
A DNA test can tell you about your genetic predisposition for or against certain traits. These traits can range dramatically, from simple information like hair or eye color to your susceptibility to mental illnesses and disabilities like schizophrenia or multiple sclerosis. Traits like these are biologically rooted, which are the only kind that a DNA test could anticipate. You might also learn that your genes indicate other interesting things, like having a high tolerance for handling stress or a tendency toward lower-than-average body temperature.
It’s important to remember that these results are only suggested by your genes, they’re not predictions or guarantees. A DNA test can’t tell you exactly what the future will hold. Many lifestyle and environmental factors can affect a gene’s activation.3
That being said, when it comes to common things you can learn about yourself from health-based DNA tests, there are many overlaps in the genes they look at. (There are only so many genes that influence your health and behavior, after all.) Some of the most common traits investigated by most — if not all — of our top picks include:
Additionally, certain tests (such as Nebula Genomics) can help you learn about your carrier status. And tests that investigate your epigenetics (like the one from MyToolbox Genomics) may be able to help you learn more about how to alter the way some of your genetics are being expressed in order to improve your health and wellness.
Are at-home DNA tests accurate?
For the most part, the kinds of DNA tests you take at home are accurate. While the accuracy depends on the kind of test you take, all of the tests we recommend operate with stringent quality control standards, such as by using CLIA- and CAP-certified laboratories.16 Laboratory standards such as CLIA (the Clinical Laboratory Improvement Amendments) are a great clue into a genetic test’s accuracy. Having CLIA certification means that the lab meets or exceeds the quality, privacy, and safety standards set by the Centers for Medicare and Medicaid Services.17
Our top three picks — Nebula Genomics, InsideTracker, and MyToolbox Genomics — also go to great lengths to back up your results with scientific research studies. You’ll often find links and citations to these studies in your results and throughout the companies’ websites.
To understand more about accuracy across the board, one of our testers gathered their results for a few overlapping traits measured by our top picks for DNA health tests. The genes for these traits are commonly analyzed in many DNA tests:
- Cilantro/coriander preference (do you think it tastes like soap?)
- Caffeine sensitivity
- Gluten intolerance
Not every test looked at all three of these genes, as indicated in the chart below.
| Cilantro preference | Caffeine sensitivity | Gluten intolerance | |
|---|---|---|---|
| Tester’s experience | Like | Normal to low | Tester has gluten intolerance |
| Nebula Genomics | Unable to tell | Low | High (many variants noted) |
| MyToolbox Genomics | Not part of test | Normal | Not part of test |
| InsideTracker | Not part of test | Normal | Average (one variant, not seen) |
You can see from these results that they were generally accurate, but none of them were perfect. Our lived experiences are as much, if not more, of a predictor than what genetic tests can tell us. This doesn’t mean that genetic tests are worthless, however — a DNA health test like the ones in this guide can help you to identify trends in your life and biological inclinations.
How can a DNA test help my health?
While a DNA test might not predict your future, it can give you a good insight into how your body works. Whether you’re driven by curiosity about your carrier status or you’re looking for a personalized exercise plan, an at-home DNA test may be able to help.
For the most part, DNA tests can help you stay vigilant about potential diseases. As an example, if you find out that you have a gene closely related to strokes, learning the four major warning signs of a stroke (the F.A.S.T. signs) can help you be proactive and prepared in ways you might not have been previously.18 These measures can range dramatically from learning symptoms and changing your diet to having preventative surgeries, such as in the case of a positive BRCA1 or BRCA2 mutation (these inherited mutations are closely linked to breast and ovarian cancers).19
We recommend familiarizing yourself with the genetic test you’re taking: What does it test for? What results might you expect? However, be aware that some genetic findings may be shocking or upsetting, so it’s important to be prepared for uncomfortable situations. Finding out that you’ve tested positive for an APOE mutation that makes you more likely to develop Alzheimer’s isn’t something you’d want to do alone.20
Genetic counselors can also help you out here, both in processing the finding and retesting to make sure that it was correct in the first place. Some companies offer this service on their platform. For instance, Sequencing.com lists genetic counseling on its app marketplace (provided by DNAVisit for $129 per genome).
What do DNA testing companies do with my information?
Every DNA testing company has a different set of rules about sharing information. Still, none of our top picks share any traceable personal data back to you without your consent. This personally identifiable information (PII) — things like your name, address, or health history — is separated from your genetic information before it goes off to the lab. DNA testing companies often have kit IDs (unique codes) that keep your information with your account but separate from any personal data to keep your information secure.
Some DNA testing companies share your genetic database with researchers (de-identified, so there’s no way to trace it back to you) to help them better understand human genetics. Sharing information like this can create more realistic ideas of how genes relate and, therefore, what they might do. However, a DNA test that shares this information needs to have informed consent from you. They might share this while you’re purchasing your kit or after creating an account.
Insider Tip: If you aren’t sure whether or not your information has been given to researchers, check the footer on the DNA testing company’s webpage. Most companies that partner with researchers have their informed consent information readily available. If there’s no information in the footer, you can also reach out to customer service and ask.
One of the tricky things about DNA testing is that it never impacts just you. Once your DNA is in a database, your family — particularly your immediate family, such as your parents, siblings, and children — has been at least partially cataloged too. This is because you all share genetic material from the same source.
Some studies have found that about 60% of all individuals of European descent have at least a third cousin’s DNA in a database, so you likely aren’t alone.21 If you’re concerned about the broader implications of having your DNA in a database like that, most testing companies are willing to delete your information from their servers if you reach out and make a request.
Nebula Genomics
Best overall, best for carrier screening, and best full genome test
Pros
- Highly-researched and fleshed-out trait analyses
- Exceptional accuracy, checking results up to 100 times
- Detailed genome exploration tools let you look at your DNA base by base
- Your results are updated weekly with new information from studies
- Extensive privacy protection methods
- Frequent discounts make testing more accessible and affordable
- Includes results for the makeup of your oral microbiome and some ancestry info
Cons
- Requires a paid membership to maintain access to your information
- No action plans or guidance provided
- A lot of scientific jargon in Explorer and Analysis tools
- Results take 12-14 weeks

Photo by Innerbody Research
Nebula Genomics is a whole-genome analysis service that provides you with a complete look at — and complete control over — your genetic information. Tools like the Genome Browser and Gene Analysis run on some of the same tools used by geneticists. Your results also include the Library, which is filled with easy-to-understand, research-backed summaries for over 340 different genetic traits. Though not as densely, MyToolbox Genomics and InsideTracker also provide extensive links and citations to scientific studies backing up their results.
Whereas the information is a bit easier for laypeople to read in the Library section, the Genome Explorer and Analysis tools are filled with scientific jargon that can sometimes be overwhelming or confusing. Because of this, we recommend using some of the time spent waiting for your results to brush up on some genetic terminology. Once you have a decent idea of what your results are talking about, then Nebula Genomics can provide you with pretty much everything you’d ever need or want to know about your genome.
We delve further into what these tools can do in our full Nebula Genomics review, but, basically the Genome Explorer lets you look at each gene in every one of your chromosomes, and Gene Analysis allows you to search your genes for any variants of note. The latter is most useful for those who are in search of specific variants that could be pathogenic. (The Human Protein Atlas and SNPedia are great resources for finding information on genes and their variants.)
However, if the thought of studying, or clicking through hundreds of traits and dense scientific reports is daunting, Nebula Genomics has a bite-sized Traits section with four categories of curated traits — appearance and hormones, behavior and perception, body and athleticism, and nutrition and diet — each exploring phenotypes (observable traits) common in many genetic tests. This information isn’t nearly as detailed or thorough as what you’d find in the Library, but it’s still interesting in its own right.

Our testers found that many of the traits in this section came with the result: “Your genotype has not been described in association with this trait.” We hope Nebula will expand on this section someday and add more genes into the mix, especially since some of the Trait section entries without associated genes have similar entries in the Library that state otherwise. For example, one of our testers received the “your genotype has not been described…” result for the height trait, but then was in the 100th percentile for height in the Library (“very high genetic predisposition to taller stature”).

And, last but certainly not least, the final two sections of the results from Nebula Genomics include your oral microbiome and ancestry.
Oral microbiome
This is a more recent addition to Nebula’s results — it wasn’t available the last time we tested its kits, but it’d been mentioned around the website, so we’re happy to finally get to see it in action.
In this section, you’ll find an interactive header filled with little cartoony, animated bacteria; this is a visualization of your oral microbiome. If you hover over a bacterium, you’ll learn its name and relative abundance in your sample. As you scroll down the page, each group of bacteria has its own entry in order of abundance (greatest to least). The information includes a description of the bacteria, how abundant it was in your sample, how your abundance compares to others’, and some studies about what that bacteria may mean for your health.
Ancestry
Nebula’s ancestry is a bit of a lukewarm experience if you don’t have a Y chromosome, and it can be confusing even if you do. If you want a more user-friendly ancestry breakdown, we recommend a service dedicated to that purpose, like AncestryDNA.
There are two Nebula ancestry reports — and if you don’t have a Y chromosome, your results will mainly be relegated to the basic one. This report using the Eurogenes K36 calculator breaks down your results by percentages of where your DNA likely came from (for example, “Italian: 5.3%”). If you do have a Y chromosome, then the deep ancestry report through YFull will provide a bit more (highly technical) information. However, be warned that the YTree is full of strings of numbers and genes that require a lot of research or specialized knowledge to truly understand.
Our Nebula Genomics testing experience

Photo by Innerbody Research
While the results may be a bit complicated at times, Nebula Genomics’ tests are very straightforward to complete. Each test kit includes:
- An instruction sheet
- Biohazard bag
- Cheek swabs
- Solution vial
- Prepaid return envelope
The instruction sheet includes a kit ID number, which you’ll need to use to register your test on the Nebula Genomics website. This ensures that your results will be linked to your account, without needing to use any personally identifying information (like your name).
After registering your kit, Nebula lists a few guidelines to follow before you can collect your sample:
- Rinse your mouth out with water 30 minutes before sample collection
- Refrain from eating, drinking, or smoking during that half-hour wait before collection
- If you’re sick, it’s recommended that you wait until you feel better before taking your sample
Once it’s time to collect your sample, you’ll rotate the swab against the inside of each cheek (one swab per cheek) for a minute, then place the swabs in the collection vial, put the vial in the biohazard bag, and ship it off in the included pre-paid envelope.
Photo by Innerbody Research
Your dashboard will let you know when your sample arrives at the lab, and results take 12-14 weeks to be ready. Our testers’ results took about 14 weeks on average. This was the longest results wait time out of any test in this guide (InsideTracker was the quickest at only ten days).
Nebula Genomics pricing
Nebula Genomics offers three different test kits, along with varying ways to purchase a membership. An active membership is required to maintain access to your test results and receive regular updates. Competitors Sequencing.com and 23andMe (only the “plus” tiers) also require memberships, but InsideTracker and MyToolbox Genomics don’t.
The chart below breaks down the various costs for Nebula Genomics.
| Kit price only | Kit + quarterly membership | Kit + yearly membership | Kit + lifetime membership | |
|---|---|---|---|---|
| Standard kit | $99 | $148.98 (Membership costs $49.98 per quarter) | $228.96 (Membership costs $129.96 per year) | $299 total |
| Deep kit | $249 | N/A | $398.88 (Membership costs $149.88 per year) | $544 total |
| Ultra Deep kit | $899 | N/A | $1,048.88 (Membership costs $149.88 per year) | $1,194 total |
If Nebula’s pricing seems a bit steep, then we recommend waiting for one of the company’s frequent sales. On major holidays and at random times throughout the year, Nebula Genomics will run promotions offering a decent amount of money off its test kits. For instance, at one point during our research of the company, there was an active sale that took the Deep test with a lifetime membership from $544 down to $374 (that’s $170 less, or about 30% off). This discounted price is only about $75 more than the DNA + Epigenetics test from MyToolbox Genomics, our best budget pick.
MyToolbox Genomics
Best budget pick and best epigenetic test
Pros
- Provides direct links to studies throughout your results
- Offers custom meal plans, a training regimen, and vitamin recommendations
- You can view results on the computer or via a mobile app
- Relatively inexpensive with no membership fees
- Checks 16 health predisposition traits
Cons
- Website and app are a bit difficult to navigate
- Sample collection can be difficult
- Frequent epigenetics testing costs can add up over time
- Some results are assumed due to trends instead of based on hard data
We think that MyToolbox Genomics is a great DNA and epigenetics health test for people on a budget. Similar to competitors InsideTracker and Nebula Genomics, the results from MyToolbox Genomics are mostly rooted in scientific research studies, and they can act as a great starting point for learning more about your DNA.
MyToolbox Genomics’ DNA test isn’t a whole genome test like Nebula, but the results are still packed with useful information. The list below breaks down what’s included in the DNA and epigenetics tests, along with the costs.
| DNA Test | Epigenetics Test | DNA + Epigenetics Test | |
|---|---|---|---|
| Cost | $199 | $273 | $299 |
| Saliva collection | |||
| Virus Risk Score | |||
| 16 Health Predisposition Traits | |||
| Custom Meal Plans | |||
| Custom Training Regimen | |||
| Vitamin Recommendations | |||
| Changeable Insights | |||
| Track DNA Expression Changes Over Time | |||
| Air Quality Impact by Location | |||
| Additional Specific-to-You Suggestions | |||
| In-App and Downloadable Results | |||
| Free In-App Updates |
Now, even though the DNA test claims to offer insight into 16 health predisposition categories, it’s actually 15 — the 16th is more of a summary than a category of its own. However, each of those 15 categories includes 5-18 individual traits; this means that MyToolbox’s DNA test checks for 115 traits in total. The following list contains each of the 15 categories.
And then the “changeable insights” of the epigenetics test measures your DNA’s epigenetic expression across five different categories:
- Biological age
- Eye age
- Hearing age
- Memory age
- Inflammation score
As we covered earlier in our guide, epigenetics is the study of how your environment, behaviors and lifestyle can lead to chemical changes that affect the way your genes work. Even your diet and exercise regimen can alter your genes’ expression.13
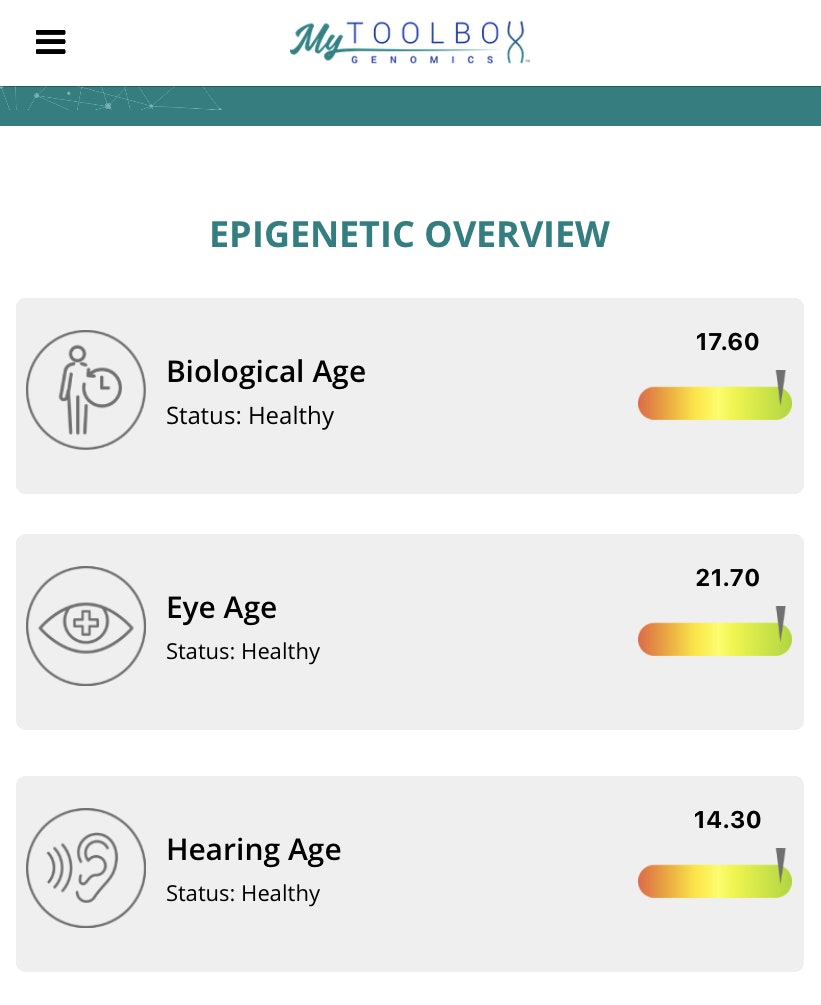
Because of these potential changes, MyToolbox Genomics recommends retesting your epigenetics every 3-12 months. You don’t necessarily need to, but it can be a good way to see how any of your lifestyle changes over time may be affecting your genes. If you do wish to retest every few months, MyToolbox offers subscription plans that reduce the cost of an epigenetic test kit, though it does come with a monthly payment.
- Retest every three months: four tests a year for $195 per test or ($65/month)
- Retest every six months: two tests a year for $204 per test ($34/month)
- Retest every 12 months: one test a year for $216 per test ($18/month)
Like InsideTracker, MyToolbox Genomics offers personalized guidance to help you improve your health based on your results. The available action plans currently include:
- Meal plans: you can choose from goals such as weight loss, fitness, muscle building, and health and well-being
- Exercise regimen: you can receive assistance in creating plans for your overall workout, warm up and cool down, injury prevention, and wellness
MyToolbox has been stating that “Lifestyle tracking” is coming soon for over a year now, but, unfortunately, all mentions of it have been removed from the website. We hope the company will consider adding more action plans in the future.
For additional information, check out our full review of MyToolbox Genomics.
Our testing experience with MyToolbox Genomics

Photo by Innerbody Research
Unlike InsideTracker and Nebula Genomics which both use cheek swabs, MyToolbox Genomics requires you to collect 4mL of saliva by spitting into a tube. Our testers found it difficult to produce enough saliva to hit the required amount, but they found it helpful to think about their favorite foods and even lemons.
As with most home testing kits, you’ll have to register your test kit using the included kit ID. You can either manually enter this code online, or scan it using the ID scanner in the MyToolbox Genomics app.
Once you send in your sample using the prepaid mailer, the wait for your results begins. If you opt for both tests, your DNA and epigenetic results will likely arrive at different times (as experienced by our testers).
- DNA results: 3-4 weeks after laboratory processing
- Epigenetic results: 5-6 weeks post-processing
While this isn’t the fastest turnaround time our testers experienced (ten days from InsideTracker), it’s still about half the time they waited for results from Nebula Genomics.
InsideTracker
Best for nutrition and fitness and fastest turnaround time
Pros
- Thorough detail and explanations of all blood biomarkers
- Interactive, engaging action plans allow for extensive personalization
- Great choice for athletes or those focused on diet, stamina, and muscle health
- Goal-setting allows for regular check-ins and monitoring without any additional fees
- Very fast results turnaround time (ten days for our testers)
- Take 20% off with coupon code INNERBODY20
Cons
- DNA testing is more of an add-on than a main focus
- One of the most expensive tests we’ve come across
- Requires retesting blood biomarkers every 3 months
- Homogenous research base means your comparisons won’t necessarily always reflect reality
InsideTracker functions more like a health monitoring service than a genetic test. In fact, its DNA testing is an add-on to its primary testing plans, which measure biomarkers in your blood for over 40 major health concerns. These results are then combined with action plans — steps you select from a list to improve your health in various categories (from cognition to fat loss and much more) based on your bloodwork results.
There are technically only two options for receiving bloodwork results from InsideTracker — the Ultimate test kit ($699) and the blood results upload subscription ($149). The latter is definitely the less expensive route to go, but you might not get as many personalized insights if your results from an outside source don’t test for all of the same things as InsideTracker. Both plans offer an analysis for the same 48 biomarkers, but you can’t get guidance for results you don’t have. You can learn more details about the 48 biomarkers in our full InsideTracker review.
But, how does your DNA come into play? InsideTracker compares your cheek swab results to your blood biomarker results to show you where you’re beating the odds and where you’re predisposed to have a harder time. The genes themselves matter less than how your bloodwork turns out, but you can still glean some interesting information about how your genetic makeup could be affecting your health. Here are the DNA insights InsideTracker offers:
- How your DNA may affect your BMI, fasting glucose, total cholesterol, LDL cholesterol, HDL cholesterol, triglycerides, and blood pressure
- Your potential to lose weight on a low-fat diet or high-saturated-fat diet
- How your genes could alter caffeine metabolism
- Your potential to need more or less sleep
- If your vitamin levels could be affected (lower magnesium, vitamin D, calcium, B12)
- Whether or not your genes could lead to lower hemoglobin, or higher white blood cell count, C-reactive protein, and GGT
- Your potential to excel at endurance activities or power activities
- If certain genes point toward higher testosterone and free testosterone levels
- If you’re genetically predisposed to gluten intolerance, lactose intolerance, or peanut allergies
Your DNA results help to provide insight into your blood biomarker results, and they can help you learn about why you may be having trouble getting certain biomarkers into their optimal ranges.
With your blood test results, you’ll receive an action plan that provides tailored advice for diet, exercise, and lifestyle changes. Some recommendations are as simple as taking a vitamin or drinking tea every day, while others may involve losing over 10lb or doing cardio exercises a few times per week.
Additionally, InsideTracker also has another blood test available, InnerAge 2.0. This test doesn’t have anything to do with your action plan, but it’s similar to MyToolbox Genomics’ biological age results (but based on biomarkers instead of epigenetics). This test’s results compare your information against others who have taken InsideTracker’s tests. For this reason, we feel that the InnerAge 2.0 results should be taken with a grain of salt — the research base is on the homogenous side. As of 2018, 85% of the InsideTracker user base is white, moderately active adults between the ages of 40 (women) and 43 (men).7 If you don’t fall into this group, your InnerAge 2.0 results likely won’t be accurate.
InsideTracker pricing
InsideTracker’s prices are some of the highest we’ve come across for this type of testing, but you can save 20% off using our coupon code INNERBODY20. The chart below breaks down the pricing of InsideTracker’s test kits and bundles.
| Price | Price with INNERBODY20 code | Cost per test (if multi-pack bundle) | |
|---|---|---|---|
| DNA Results Upload | $29 | $23.20 | |
| DNA Kit | $249 | $199.20 | |
| Blood Results Upload Subscription | $149 | $119.20 | |
| Ultimate | $699 | $559.20 | |
| 2 Ultimate Bundle | $1,388 | $1,110.40 | $694 ($555.20 with code) |
| 4 Ultimate Bundle | $2,682 | $2,145.60 | $670.50 ($536.40 with code) |
| Ultimate + DNA + InnerAge Bundle | $921 | $736.80 |
It’s worth mentioning that InsideTracker recommends retesting your blood biomarkers every three months — this can make costs add up fast. If you’re interested in frequent retesting, then opting for a bundle and using our coupon code should save you a fair bit of money versus purchasing individual Ultimate tests.
Sequencing.com
Pros
- Upload your DNA results from other major companies for free
- Wide range of apps in the marketplace to analyze your DNA
- Results come within 24 hours from individual marketplace apps
- Whole genome testing
- Strong internal privacy measures
Cons
- Uploads using outside data don’t always work; your info may be deemed “incompatible”
- Uploading full genomic data takes over six hours, even with very fast internet speeds
- Full genome sequencing takes three months to complete
- Additional fees and membership costs after your test
- Third-party apps have unclear privacy policies
- Website is not always intuitive to navigate
On paper, Sequencing.com sounds like a phenomenal option. It’s an app-based marketplace for genetic testing, where you can either upload genetic results from any major DNA testing company for free or have Sequencing.com perform full genome testing for a (somewhat expensive) fee. Once you have your genetic results, you can pick from more than 50 investigative third-party testing apps to personalize what you learn about your genetic code.
Since many of these apps are third-party, Sequencing.com did not create or run the tests. In our testing, we found the quality of these third-party tests varied dramatically. Some provided almost 600 pages of dense information, while others gave a few words on a handful of pages. Some reports cost upwards of $60, giving the impression that they’d be high quality while turning over difficult-to-read or surface-level information.
On top of having to purchase additional reports that you’d have included with a competing service like Nebula Genomics, you also need to subscribe to a Sequencing.com membership. At least with Nebula, your membership includes additional reports for free.
Here’s how the pricing breaks down for Sequencing.com (all tests include whole genome sequencing):
| Cost | Includes | |
|---|---|---|
| Wellness Screen | $399 | One report (Wellness and Longevity) |
| Ehlers Danlos Screen | $399 | One report (Next-Gen Disease Screen) |
| Women’s Health Disease Screen | $449 | Four reports (Next-Gen Disease Screen; Prevent Breast Cancer Report; Melanoma Skin Cancer Report; Healthcare Professional Report) |
| Comprehensive Health Screen | $499 | Five reports (Healthcare Professional Report; Next-Gen Disease Screen; Wellness and Longevity; Prevent Breast Cancer Report; Melanoma Skin Cancer Report) |
| 4 Week Turnaround | $1,399 | Five reports (Healthcare Professional Report; Next-Gen Disease Screen; Wellness and Longevity; Prevent Breast Cancer Report; Melanoma Skin Cancer Report) |
| 2-3 Week Turnaround | $1,999 | Six reports (Healthcare Professional Report; Next-Gen Disease Screen; Wellness and Longevity; Prevent Breast Cancer Report; Melanoma Skin Cancer Report; Medication and Drug Response) |
As you can see, the test kits from Sequencing.com are pretty pricey compared to competitors. And this is before you take into consideration the need to purchase a membership and additional reports from the marketplace.
There are three paid membership options available for Sequencing.com:
- Silver ($23.98/month): get two free apps from the marketplace every month
- Gold ($69.98/month): get four free apps every month and real-time health updates
- Platinum ($119.98/month): get six free apps per month, real-time health updates, and lifetime live genetic counseling
Sequencing.com offers an interesting approach to DNA testing, but we can’t recommend it as our top choice at this time due to how expensive it is compared to other options that provide a lot more information at a much lower price. For example, a Comprehensive Health Screen test combined with the Gold level membership will cost a total of $568.98 — that’s more than a Deep kit with a lifetime membership from Nebula Genomics ($544). And you’ll need to keep paying almost $70 each month to maintain your Sequencing.com membership, there are no lifetime options.
23andMe
Pros
- Only FDA-approved at-home genetic test
- Combines ancestry and health DNA while still giving thorough results in both
- Website is user-friendly, with easy access to your results
- Provides simple, yet accurate explanations and points out any concerns
- Basic Ancestry and Ancestry + Health tests are relatively affordable
Cons
- Recent data breach compromised sensitive information of 6.9 million clients
- Exome testing option is more expensive than full genome tests on the market
- Membership required for 23andMe+ tiers and to receive updates on lower tiers
- A lot of surveys
If you’ve heard of just one genetic test, it’s likely to be 23andMe. This popular DNA testing service is currently the only one approved by the FDA,12 meaning it’s the only one that can legally provide medical advice, such as if it notifies you of a carrier status or alarming variations on genes like BRCA1 (which increases your risk of breast and ovarian cancer by up to 72%).22
However, FDA approval for genetic tests is slow since a test for each gene must be approved separately. This process is painfully slow for a service like Nebula Genomics, which looks at all 20,000-plus genes in the human genome.
Insider Tip: 23andMe recently experienced a massive hack that compromised the sensitive ancestry (and some health) data of nearly seven million clients.9 Until the company improves its security measures (and stops blaming breach victims for its own shortcomings) we recommend opting for another DNA health test that strives to protect your data — like any one of our top three picks (Nebula Genomics, MyToolbox Genomics, and InsideTracker).23
23andMe’s testing spans a wide range of categories, giving you a small sample of everything from your ancestry to carrier status to pharmacogenetics (how you’re genetically inclined to react to different drugs). It samples specific genes and adds new traits regularly, but updates don’t always provide more insight into your genes — more often than not, they are newly added for those who are just getting started with 23andMe. A paid membership allows you to access these updates.
Recently, 23andMe has added exome sequencing options. Exome tests look at your DNA’s exons (the parts of your DNA that provide instructions for making proteins). The exome only makes up about 1% of your genome, but it can still be an efficient method for finding mutations. However, we find it odd that 23andMe charges nearly the same amount to decode 1% of your DNA ($1,188 per year) as Nebula Genomics does to analyze your whole genome with the Ultra Deep kit and a lifetime membership ($1,194 total, no recurring charges).
The following list details the costs of each version of 23andMe:
- Ancestry Service: $109
- Health + Ancestry Service: $129
- 23andMe+ Premium: $159 (renews at $69 per year)
- 23andMe+ Total Health: $99/month (billed as one payment of $1,188 per year)
Any 23andMe kit that examines your health (so, all of them but the basic ancestry test) checks for a decent variety of traits, including:
- Physical traits: things like flat feet, motion sickness, toe length, and eye color
- Carrier status: if you carry, but don’t experience symptoms of genetic illnesses that can be passed to offspring
- Health predispositions (only with 23andMe+): gestational diabetes, anxiety, and eczema
- Wellness (only with 23andMe+): pet allergies, lactose intolerance, and caffeine sensitivity
- Pharmacogenetics (only with 23andMe+): how you metabolize medications
- Exome sequencing (only with 23andMe+ Total Health): checks if you have a hereditary predispositions to health concerns like cancer, cardiovascular disease, kidney disease, and others
- Blood test panels (only with 23andMe+ Total Health): includes a comprehensive metabolic panel, CBC, lipid panel, and endocrine system blood tests
Like Nebula Genomics and InsideTracker, 23andMe has partnered with researchers to try and learn more about the human genome, but the company needs data to do so. As with competitors, participating in 23andMe Research is optional, but you’ll have to navigate away from some surveys and redact your consent to keep away your de-identified and anonymized data. It’s important to note that 23andMe links three informed consent documents to its research page:
- Research consent
- Biobanking consent
- Individual data-sharing consent
You’ll need to give your consent — or withdraw it — at the start of the process. If you ever forget exactly what you’ve consented to, 23andMe makes it easy to find and recall.
FAQ about DNA testing
Sources
Innerbody uses only high-quality sources, including peer-reviewed studies, to support the facts within our articles. Read our editorial process to learn more about how we fact-check and keep our content accurate, reliable, and trustworthy.
Rogers, K. (n.d.). Genographic Project. Britannica.
National Institutes of Health. (2022). First complete sequence of a human genome. NIH Research Matters.
National Library of Medicine. (n.d.). What does it mean to have a genetic predisposition to a disease? MedlinePlus.
Qin, D. (2019). Next-generation sequencing and its clinical application. Cancer Biology & Medicine, 16(1), 4-10.
Salk, J. J., Schmitt, M. W., & Loeb, L. A. (2018). Enhancing the accuracy of next-generation sequencing for detecting rare and subclonal mutations. Nature Reviews Genetics, 19(5), 269-285.
Illumina. (n.d.). Experiments & Protocols: Coverage depth recommendations. Illumina, Inc.
Westerman, K., Reaver, A., Roy, C., Ploch, M., Sharoni, E., Nogal, B., Sinclair, D. A., Katz, D. L., Blumberg, J. B., & Blander, G. (2018). Longitudinal analysis of biomarker data from a personalized nutrition platform in healthy subjects. Scientific Reports, 8(1), 1-10.
Wetterstrand, K. (2021). The Cost of Sequencing a Human Genome. NIH National Human Genome Research Institute.
Brown, K. (2023). 23andMe Hack Breaches 6.9 Million Users’ Info, Including Some’s Health Data. Time.
Jayson, S. (2023). Waiting for Medical Test Results: Dealing With the Anxiety, Uncertainty. AARP.
Hanchard, N. (2024). Carrier. NIH National Human Genome Research Institute.
U.S. Food and Drug Administration. (2017). FDA allows marketing of first direct-to-consumer tests that provide genetic risk information for certain conditions. FDA.
Centers for Disease Control and Prevention. (2024). Epigenetics, Health, and Disease. CDC.
Fang, F., Andersen, A. M., Philibert, R., & Hancock, D. B. (2023). Epigenetic biomarkers for smoking cessation. Addiction Neuroscience, 6.
Yu, M., Hazelton, W. D., Luebeck, G. E., & Grady, W. M. (2020). Epigenetic aging: More than just a clock when it comes to cancer. Cancer Research, 80(3), 367.
College of American Pathologists. (n.d.). Laboratory Accreditation Program. CAP.
U.S. Centers for Medicare & Medicaid Services. (2023). Clinical Laboratory Improvement Amendments (CLIA). CMS.
American Stroke Association. (2021). Stroke Symptoms. American Heart Association, Inc.
Centers for Disease Control and Prevention. (2023). BRCA1 and BRCA2. CDC.
National Institute on Aging. (2021). Study reveals how APOE4 gene may increase risk for dementia. NIH.
Erlich, Y., Shor, T., & Carmi, S. (2018). Identity inference of genomic data using long-range familial searches. Science, 362(6415), 690-694.
National Cancer Institute. (2020). BRCA Gene Mutations: Cancer Risk and Genetic Testing. NIH.
Arntz, P. (2024). 23andMe blames “negligent” breach victims, says it’s their own fault. Malwarebytes Cyber Security Software.







